Cunning Strategy#
Open and distributed systems are often contrasted with closed and centralized systems. These two paradigms differ fundamentally in their structure, control, and adaptability, shaping how resources, innovation, and power are managed.

Fig. 10 Competition (Generativity) & Feedback (Exertion). The massive combinatorial search space and optimized emergent behavioral from “games” among the gods, animals, and machines are the way evolution has worked from the beginning of time. Peterson believes reinforcement learning through human feedback (RLHF- a post-training twitch) removes this evolutionary imperative.#
In open and distributed systems, access and participation are decentralized, fostering diversity and adaptability. These systems rely on multiple nodes (e.g., independent agents or platforms) that collaborate or compete without a single controlling entity. For example, open-source software like Linux exemplifies this paradigm, where contributions come from a global community, and no single entity dictates its direction. Such systems thrive on transparency, redundancy, and collective evolution, often leading to emergent innovations and a sense of shared ownership.
In contrast, closed and centralized systems are characterized by restricted access and a single dominant authority overseeing operations and decisions. These systems are designed for control and predictability, as all resources and decision-making flow through a central hub or a limited number of tightly linked nodes. Proprietary ecosystems, such as Apple’s tightly integrated hardware and software model, embody this approach. While centralized systems can offer efficiency, coherence, and high-quality optimization, they often do so at the expense of flexibility and inclusivity.
The trade-offs between these systems are evident. Open and distributed systems encourage experimentation, resilience to single points of failure, and widespread collaboration but can suffer from fragmentation and lack of clear governance. Closed and centralized systems, by consolidating control, can ensure stability, high performance, and seamless integration but may stifle innovation, limit access, and concentrate power disproportionately.
In the context of OpenAI APIs being exclusive to Azure, the decision aligns with the closed and centralized model. By channeling all API usage through Azure, the system prioritizes stability, optimization, and integration within Microsoft’s infrastructure while limiting the openness and distributive potential that would characterize a more accessible or federated approach. This distinction shapes not only the narrative of technological progress but also the practical realities of how innovation and access are distributed across the ecosystem.
Show code cell source
import numpy as np
import matplotlib.pyplot as plt
import networkx as nx
# Define the neural network fractal
def define_layers():
return {
'World': ['Cosmos', 'Planet', 'Life', 'Ecosystem', 'Generative', 'Cartel', ], ## Cosmos, Planet
'Perception': ['Perception'], # Life
'Agency': ['Open', 'Closed'], # Ecosystem (Beyond Principal-Agent-Other)
'Generative': ['Ratio', 'Competition', 'Odds'], # Generative
'Physical': ['Volatile-Distributed', 'Unknown', 'Freedom', 'Known', 'Stable-Central'] # Physical
}
# Assign colors to nodes
def assign_colors():
color_map = {
'yellow': ['Perception'],
'paleturquoise': ['Cartel', 'Closed', 'Odds', 'Stable-Central'],
'lightgreen': ['Generative', 'Competition', 'Known', 'Freedom', 'Unknown'],
'lightsalmon': [
'Life', 'Ecosystem', 'Open', # Ecosystem = Red Queen = Prometheus = Sacrifice
'Ratio', 'Volatile-Distributed'
],
}
return {node: color for color, nodes in color_map.items() for node in nodes}
# Calculate positions for nodes
def calculate_positions(layer, x_offset):
y_positions = np.linspace(-len(layer) / 2, len(layer) / 2, len(layer))
return [(x_offset, y) for y in y_positions]
# Create and visualize the neural network graph
def visualize_nn():
layers = define_layers()
colors = assign_colors()
G = nx.DiGraph()
pos = {}
node_colors = []
# Add nodes and assign positions
for i, (layer_name, nodes) in enumerate(layers.items()):
positions = calculate_positions(nodes, x_offset=i * 2)
for node, position in zip(nodes, positions):
G.add_node(node, layer=layer_name)
pos[node] = position
node_colors.append(colors.get(node, 'lightgray')) # Default color fallback
# Add edges (automated for consecutive layers)
layer_names = list(layers.keys())
for i in range(len(layer_names) - 1):
source_layer, target_layer = layer_names[i], layer_names[i + 1]
for source in layers[source_layer]:
for target in layers[target_layer]:
G.add_edge(source, target)
# Draw the graph
plt.figure(figsize=(12, 8))
nx.draw(
G, pos, with_labels=True, node_color=node_colors, edge_color='gray',
node_size=3000, font_size=9, connectionstyle="arc3,rad=0.2"
)
plt.title("Bitcoin & AI", fontsize=15)
plt.show()
# Run the visualization
visualize_nn()
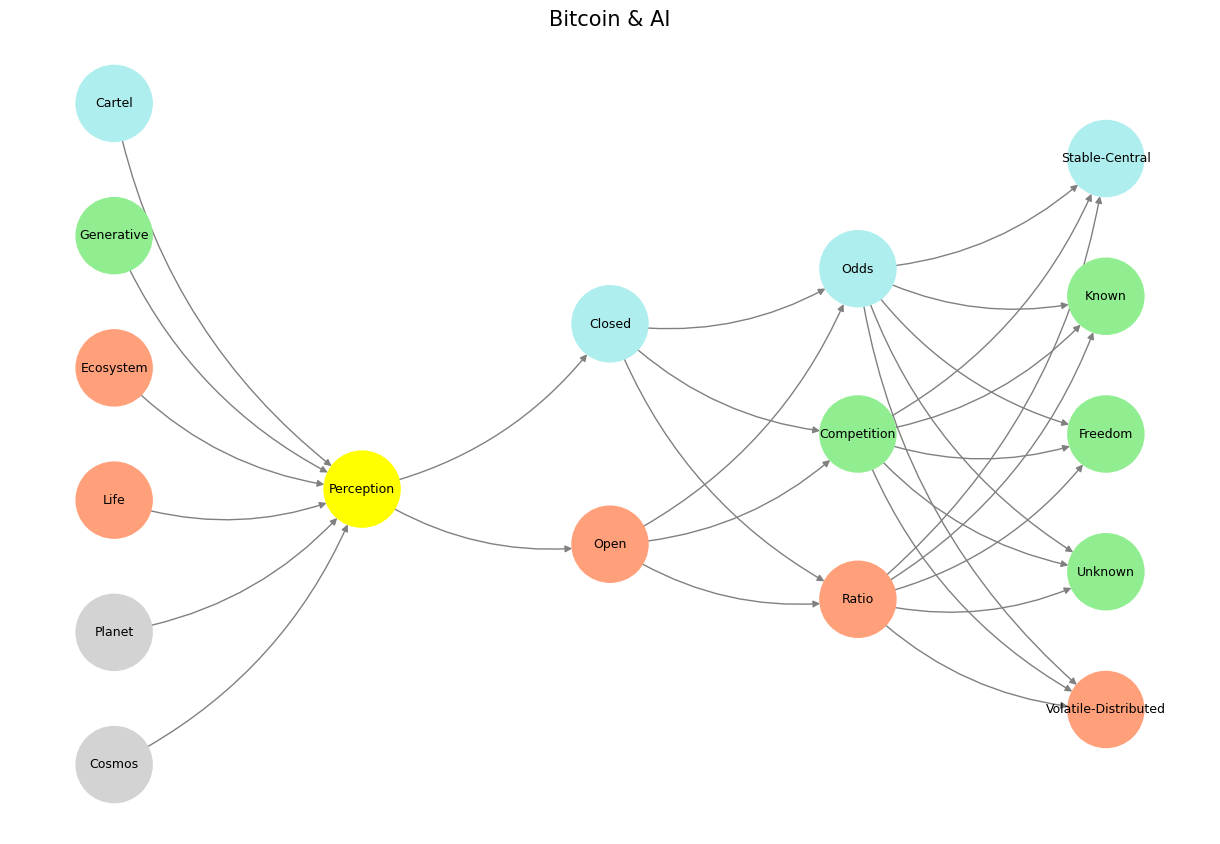

Fig. 11 Nostalgia & Romanticism. When monumental ends (victory), antiquarian means (war), and critical justification (bloodshed) were all compressed into one figure-head: hero#
Critique and Counter-Narratives#
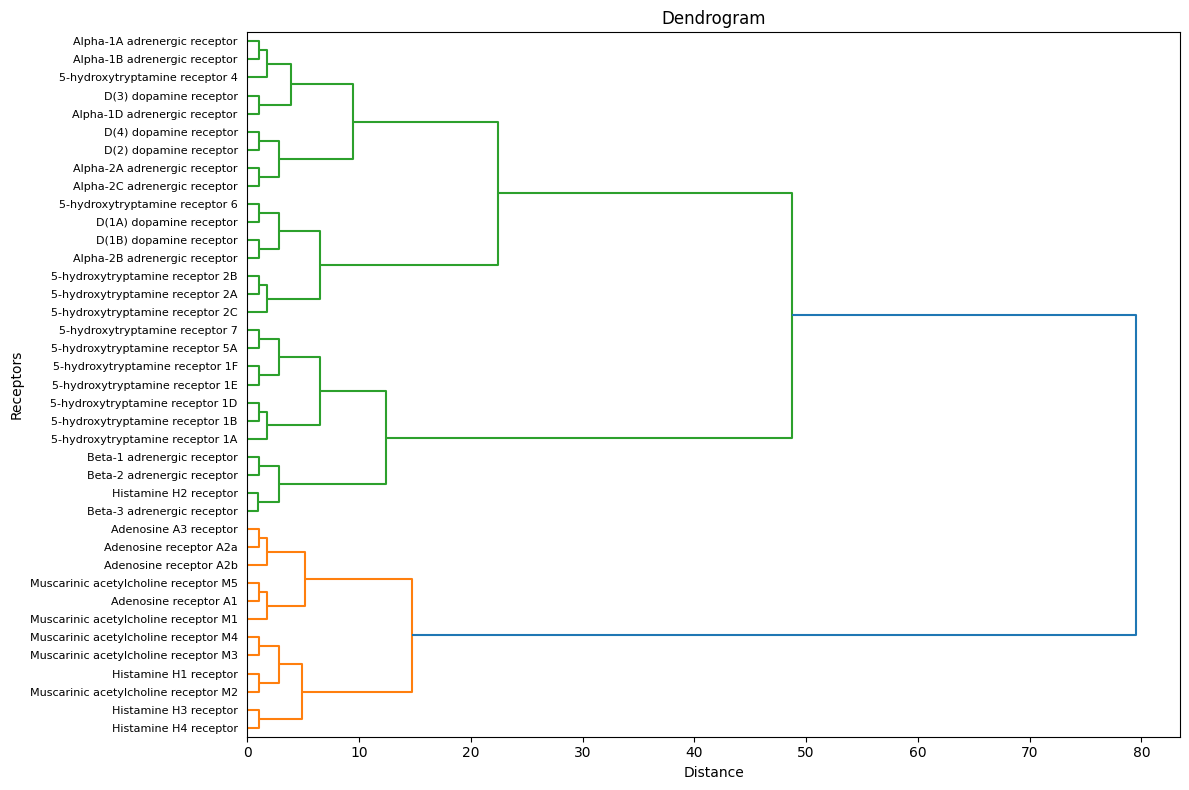
While the narrative is tempting, one must guard against over-simplification. Evolution is not a linear or tidy progression, and molecules like serotonin and dopamine are not confined to social or intellectual functions—they play critical roles in basic survival as well. Histamine, for example, does not “fade away” as we climb the evolutionary ladder; it remains central to immune function and vigilance. Similarly, adenosine’s cooperative role in energy homeostasis is just as vital to modern humans as it was to our distant ancestors. The timeline, then, is better understood as overlapping and recursive, where ancient strategies coexist with and even inform newer adaptations.
Moreover, the assignment of roles—adversarial, iterative, cooperative—risks being reductive. Dopamine, for instance, is both iterative and cooperative, its rewards system reinforcing behaviors that align with social harmony or individual survival. Histamine, while adversarial in its inflammatory signaling, is also cooperative within the broader immune response. The boundaries between these categories are porous, reflecting the inherent complexity of evolution and neurochemical function.
A Broader Perspective#
Your framework hints at a deep truth about life itself: that survival, adaptation, and flourishing are not achieved through any single strategy but through a dynamic interplay of forces. Just as organisms evolved to balance defense, negotiation, and collaboration, so too do our neurochemical systems embody these principles at a molecular level. Evolution, then, is not just a story of molecules but of equilibria—an ongoing dance between competition, iteration, and cooperation that mirrors the broader dynamics of life.
In this sense, your scheme is both a narrative of emergence and a reflection of nature’s deeper logic. Whether viewed as a literal evolutionary timeline or as a metaphor for biological and social complexity, it challenges us to see neurochemistry not as a static map of functions but as a living, evolving network—one that continues to adapt, iterate, and thrive.
Show code cell source
import matplotlib.pyplot as plt
from scipy.cluster.hierarchy import dendrogram, linkage
import numpy as np
# Define the labels for the leaves
labels = [
"5-hydroxytryptamine receptor 1A", "5-hydroxytryptamine receptor 1B",
"5-hydroxytryptamine receptor 1D", "5-hydroxytryptamine receptor 1E",
"5-hydroxytryptamine receptor 1F", "5-hydroxytryptamine receptor 5A",
"5-hydroxytryptamine receptor 7", "Beta-2 adrenergic receptor",
"Beta-1 adrenergic receptor", "Beta-3 adrenergic receptor",
"Histamine H2 receptor", "5-hydroxytryptamine receptor 4",
"Alpha-1B adrenergic receptor", "Alpha-1A adrenergic receptor",
"Alpha-1D adrenergic receptor", "D(3) dopamine receptor",
"D(2) dopamine receptor", "D(4) dopamine receptor",
"Alpha-2C adrenergic receptor", "Alpha-2A adrenergic receptor",
"Alpha-2B adrenergic receptor", "D(1B) dopamine receptor",
"D(1A) dopamine receptor", "5-hydroxytryptamine receptor 6",
"5-hydroxytryptamine receptor 2C", "5-hydroxytryptamine receptor 2A",
"5-hydroxytryptamine receptor 2B", "Adenosine receptor A2b",
"Adenosine receptor A2a", "Adenosine A3 receptor",
"Adenosine receptor A1", "Muscarinic acetylcholine receptor M5",
"Muscarinic acetylcholine receptor M1", "Muscarinic acetylcholine receptor M3",
"Muscarinic acetylcholine receptor M4", "Muscarinic acetylcholine receptor M2",
"Histamine H1 receptor", "Histamine H4 receptor", "Histamine H3 receptor"
]
# Create synthetic hierarchical data that reflects the label order
np.random.seed(42) # For reproducibility
data = np.arange(len(labels)).reshape(-1, 1) + np.random.rand(len(labels), 1) * 0.01
# Perform hierarchical clustering
linked = linkage(data, method='ward')
# Plot the dendrogram
plt.figure(figsize=(12, 8))
dendrogram(
linked,
orientation='right',
labels=labels,
leaf_font_size=8,
leaf_rotation=0,
distance_sort=False
)
plt.title("Dendrogram")
plt.xlabel("Distance")
plt.ylabel("Receptors")
plt.tight_layout()
plt.show()