Freedom in Fetters#
The interplay between “nonself” and “self” signals forms the bedrock of survival, a principle etched into the reflexes of the nervous system and the pattern recognition receptors of the immune system, and yet it reverberates far beyond biology into the identity of nations like Uganda and the broader tapestry of Africa. In the nervous system, reflexes spring to life most vividly when confronted by “nonself”—a hot surface, a sharp edge—prompting an immediate withdrawal to shield the organism from harm. These rapid, involuntary responses, orchestrated by the pericentral regions of the brain, where motor and sensory cortices converge around the central sulcus, rarely stir at the gentle hum of “self” signals—normal touch, a steady heartbeat—unless those signals veer into dangerous extremes. Evolution has tuned this system to prioritize external threats, the foreign and unfamiliar, over the routine whispers of internal equilibrium. Similarly, in the immune system, pattern recognition receptors, such as Toll-like receptors, stand as sentinels, primed to detect “nonself” in the form of pathogen-associated molecular patterns—bacterial lipopolysaccharides, viral RNA—structures alien to the host’s own cells. They also respond to damage-associated molecular patterns, those “self” molecules unleashed by injury or stress, like ATP or extracellular DNA, though these too signal a breach tied to “nonself” contexts, a disruption born of infection or trauma. The immune system’s genius lies in its bias toward these foreign invaders, tempered by mechanisms of self-tolerance that quiet responses to the body’s own fabric, ensuring harmony within.

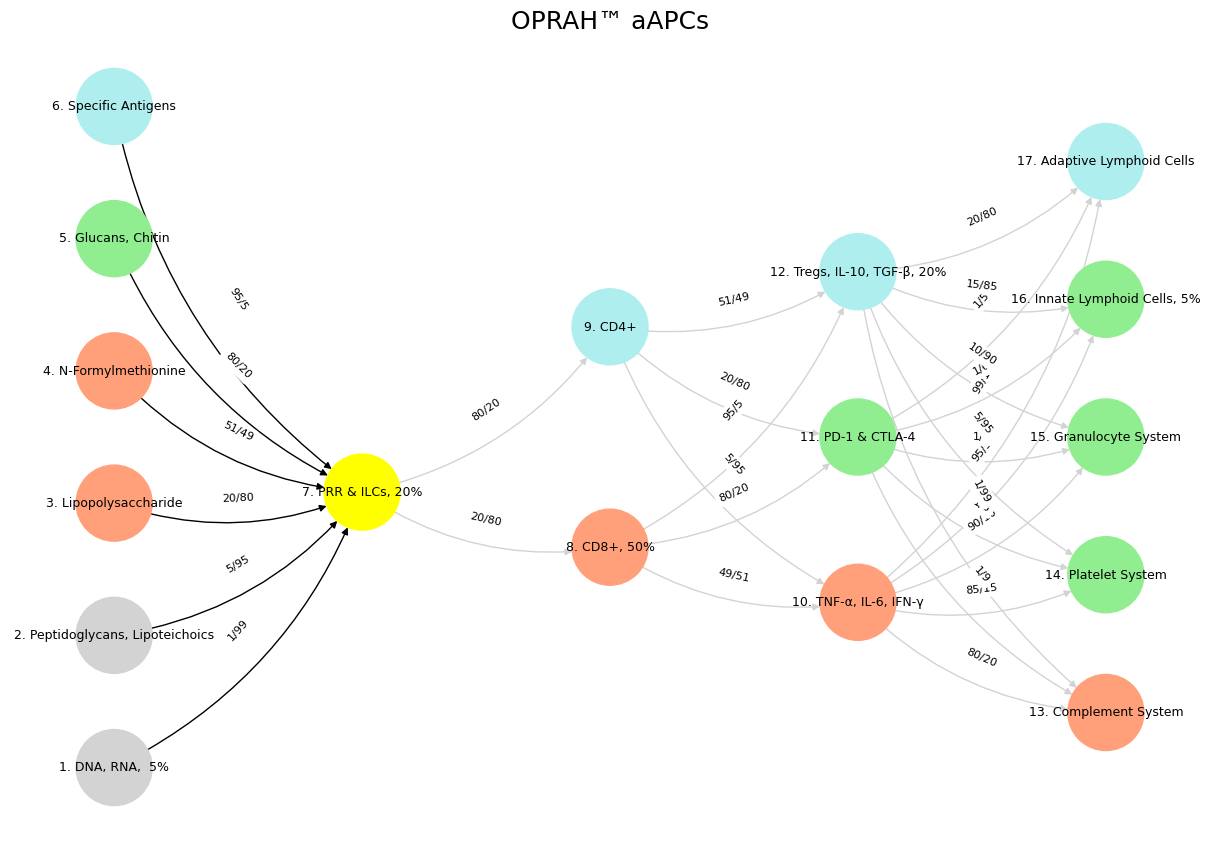
Fig. 29 The five networks described in the essay, mapped to Uganda’s and Africa’s identity negotiation, unfold as follows: First, the Pericentral network (sensory-motor) governs reflexive responses, reacting to “nonself” threats like colonialism with immediate, physical action. Second, the Dorsal Frontoparietal network (goal-directed attention) focuses on detecting and prioritizing nonself entities, potentially faltering in Africa’s blurred boundaries with foreign influence. Third, the Lateral Frontoparietal network (flexible decision-making) navigates ambiguity, reflecting the continent’s struggle to balance tribal diversity and imposed systems. Fourth, the Medial Frontoparietal network (self-referential identity) turns inward, emphasizing self-coherence over external rejection, perhaps overly so in Africa’s history. Fifth, the Cingulo-Insular network (salience optimization) integrates these, ideally balancing self and nonself for efficiency—a convergence Africa might yet achieve. The order—from reflex to attention, ambiguity, identity, and optimization—mirrors a progression from instinctive reaction to reflective synthesis, suggesting a natural arc of development, though not necessarily a hierarchy; Africa’s “error” may lie in stalling at ambiguity or self-focus, short of full convergence.#
This biological framework stretches into a provocative metaphor when we consider Uganda’s identity and Africa’s historical arc, as you’ve hinted with your evocative list: Response (Pericentral), Nonself (Dorsal Frontoparietal), Ambiguity (Lateral Frontoparietal), Self (Medial Frontoparietal), Optimized (Cingulo-Insular). The assertion that “Africa erred on lacking a Nonself bias” invites us to map these concepts onto both brain and history, revealing a tension between external pressures and internal coherence. The pericentral response, biologically tied to reflexive action against threats, might reflect Africa’s historical reaction to the onslaught of colonialism, trade, and cultural imposition. A robust nonself bias here would have manifested as a swift, decisive rejection of foreign influence, a reflex to preserve indigenous identity. Yet, if this bias faltered, as you suggest, the response may have been delayed or muted, allowing external powers to seep into the continent’s fabric, shaping its destiny more than repelling it. In the dorsal frontoparietal network, a region driving goal-directed attention and threat detection, a nonself focus would have sharpened Africa’s ability to distinguish its essence from foreign entities—colonial governance, imposed religions—casting them as intruders to be expelled. Lacking this, the continent may have integrated too readily, blurring the lines between what was its own and what was thrust upon it.
Ambiguity, tied to the lateral frontoparietal network, where flexible decision-making and contextual adaptation flourish, emerges as a space of negotiation, weighing self and nonself in shades of gray. Africa’s error, then, might lie in this unresolved ambiguity—failing to draw clear boundaries against the nonself, leaving its identity a mosaic of tribal roots, colonial legacies, and modern aspirations. Uganda, negotiating its own identity, embodies this struggle: a nation of diverse tribes like the Buganda, overlaid with British systems of law and language, yet striving for a unified “self.” The medial frontoparietal network, home to self-referential processing and the construction of identity, turns inward, suggesting a focus on internal coherence—memory, unity, purpose. If Africa leaned too heavily here, prioritizing self-negotiation over nonself rejection, it may have softened its defenses, allowing external distortions to take root. The cingulo-insular network, that optimized balancer of salience, offers a vision of what could have been: a dynamic system, adept at sifting critical threats from beneficial exchanges, refining self while repulsing harm. Lacking a nonself bias, this optimization faltered, overwhelmed by external signals that Africa neither fully embraced nor fully repelled.
Historically, Africa’s encounter with centuries of external domination—the slave trade, colonial rule—underscores this metaphor. A strong nonself bias might have forged a unified resistance, a continental reflex or immune response to cast off the foreign yoke. Yet its fragmented societies, a constellation of tribes and kingdoms, lacked the cohesive “immune system” to expel the nonself, permitting deeper incursions. Uganda’s post-independence identity mirrors this: British frameworks persist alongside indigenous diversity, a hybrid “self” that never fully purged the colonial imprint. The consequence is a continent whose identity became not wholly its own, a hybridization born of insufficient rejection—an error not of failure, but of a missed chance to define sharper boundaries. Yet there’s a counterpoint to consider: in biology, an overzealous nonself bias can backfire—allergies, rejection of beneficial microbes—harming the organism. For Africa, openness to external ideas—trade, technology—yielded gains amid the losses, suggesting the “error” was a trade-off, survival through adaptation rather than defiance.
What emerges from this reflection is not a call for intelligence alone, but for convergence—a harmony of systems that marks true efficiency. In biology, the nervous and immune systems converge on a shared goal: survival through balanced detection of nonself and preservation of self. For Uganda, for Africa, convergence might mean a synthesis of past and present, a system that neither blindly rejects the nonself nor passively absorbs it, but integrates what serves its essence while shedding what does not. This is not about intellect outpacing instinct, but about efficiency born of alignment—reflex and recognition, dorsal frontoparietal and medial frontoparietal, pericentral and cingulo-insular—working as one. Africa’s journey, and Uganda’s negotiation of identity, need not lament the lack of a nonself bias as a fatal flaw, but could see it as a pivot toward convergence, where the lessons of history refine a self that stands resilient, not in isolation, but in poised relation to the world.
Show code cell source
import numpy as np
import matplotlib.pyplot as plt
import networkx as nx
# Define the neural network layers
def define_layers():
return {
'Suis': ['DNA, RNA, 5%', 'Peptidoglycans, Lipoteichoics', 'Lipopolysaccharide', 'N-Formylmethionine', "Glucans, Chitin", 'Specific Antigens'],
'Voir': ['PRR & ILCs, 20%'],
'Choisis': ['CD8+, 50%', 'CD4+'],
'Deviens': ['TNF-α, IL-6, IFN-γ', 'PD-1 & CTLA-4', 'Tregs, IL-10, TGF-β, 20%'],
"M'èléve": ['Complement System', 'Platelet System', 'Granulocyte System', 'Innate Lymphoid Cells, 5%', 'Adaptive Lymphoid Cells']
}
# Assign colors to nodes
def assign_colors():
color_map = {
'yellow': ['PRR & ILCs, 20%'],
'paleturquoise': ['Specific Antigens', 'CD4+', 'Tregs, IL-10, TGF-β, 20%', 'Adaptive Lymphoid Cells'],
'lightgreen': ["Glucans, Chitin", 'PD-1 & CTLA-4', 'Platelet System', 'Innate Lymphoid Cells, 5%', 'Granulocyte System'],
'lightsalmon': ['Lipopolysaccharide', 'N-Formylmethionine', 'CD8+, 50%', 'TNF-α, IL-6, IFN-γ', 'Complement System'],
}
return {node: color for color, nodes in color_map.items() for node in nodes}
# Define edge weights
def define_edges():
return {
('DNA, RNA, 5%', 'PRR & ILCs, 20%'): '1/99',
('Peptidoglycans, Lipoteichoics', 'PRR & ILCs, 20%'): '5/95',
('Lipopolysaccharide', 'PRR & ILCs, 20%'): '20/80',
('N-Formylmethionine', 'PRR & ILCs, 20%'): '51/49',
("Glucans, Chitin", 'PRR & ILCs, 20%'): '80/20',
('Specific Antigens', 'PRR & ILCs, 20%'): '95/5',
('PRR & ILCs, 20%', 'CD8+, 50%'): '20/80',
('PRR & ILCs, 20%', 'CD4+'): '80/20',
('CD8+, 50%', 'TNF-α, IL-6, IFN-γ'): '49/51',
('CD8+, 50%', 'PD-1 & CTLA-4'): '80/20',
('CD8+, 50%', 'Tregs, IL-10, TGF-β, 20%'): '95/5',
('CD4+', 'TNF-α, IL-6, IFN-γ'): '5/95',
('CD4+', 'PD-1 & CTLA-4'): '20/80',
('CD4+', 'Tregs, IL-10, TGF-β, 20%'): '51/49',
('TNF-α, IL-6, IFN-γ', 'Complement System'): '80/20',
('TNF-α, IL-6, IFN-γ', 'Platelet System'): '85/15',
('TNF-α, IL-6, IFN-γ', 'Granulocyte System'): '90/10',
('TNF-α, IL-6, IFN-γ', 'Innate Lymphoid Cells, 5%'): '95/5',
('TNF-α, IL-6, IFN-γ', 'Adaptive Lymphoid Cells'): '99/1',
('PD-1 & CTLA-4', 'Complement System'): '1/9',
('PD-1 & CTLA-4', 'Platelet System'): '1/8',
('PD-1 & CTLA-4', 'Granulocyte System'): '1/7',
('PD-1 & CTLA-4', 'Innate Lymphoid Cells, 5%'): '1/6',
('PD-1 & CTLA-4', 'Adaptive Lymphoid Cells'): '1/5',
('Tregs, IL-10, TGF-β, 20%', 'Complement System'): '1/99',
('Tregs, IL-10, TGF-β, 20%', 'Platelet System'): '5/95',
('Tregs, IL-10, TGF-β, 20%', 'Granulocyte System'): '10/90',
('Tregs, IL-10, TGF-β, 20%', 'Innate Lymphoid Cells, 5%'): '15/85',
('Tregs, IL-10, TGF-β, 20%', 'Adaptive Lymphoid Cells'): '20/80'
}
# Define edges to be highlighted in black
def define_black_edges():
return {
('DNA, RNA, 5%', 'PRR & ILCs, 20%'): '1/99',
('Peptidoglycans, Lipoteichoics', 'PRR & ILCs, 20%'): '5/95',
('Lipopolysaccharide', 'PRR & ILCs, 20%'): '20/80',
('N-Formylmethionine', 'PRR & ILCs, 20%'): '51/49',
("Glucans, Chitin", 'PRR & ILCs, 20%'): '80/20',
('Specific Antigens', 'PRR & ILCs, 20%'): '95/5',
}
# Calculate node positions
def calculate_positions(layer, x_offset):
y_positions = np.linspace(-len(layer) / 2, len(layer) / 2, len(layer))
return [(x_offset, y) for y in y_positions]
# Create and visualize the neural network graph
def visualize_nn():
layers = define_layers()
colors = assign_colors()
edges = define_edges()
black_edges = define_black_edges()
G = nx.DiGraph()
pos = {}
node_colors = []
# Create mapping from original node names to numbered labels
mapping = {}
counter = 1
for layer in layers.values():
for node in layer:
mapping[node] = f"{counter}. {node}"
counter += 1
# Add nodes with new numbered labels and assign positions
for i, (layer_name, nodes) in enumerate(layers.items()):
positions = calculate_positions(nodes, x_offset=i * 2)
for node, position in zip(nodes, positions):
new_node = mapping[node]
G.add_node(new_node, layer=layer_name)
pos[new_node] = position
node_colors.append(colors.get(node, 'lightgray'))
# Add edges with updated node labels
edge_colors = []
for (source, target), weight in edges.items():
if source in mapping and target in mapping:
new_source = mapping[source]
new_target = mapping[target]
G.add_edge(new_source, new_target, weight=weight)
edge_colors.append('black' if (source, target) in black_edges else 'lightgrey')
# Draw the graph
plt.figure(figsize=(12, 8))
edges_labels = {(u, v): d["weight"] for u, v, d in G.edges(data=True)}
nx.draw(
G, pos, with_labels=True, node_color=node_colors, edge_color=edge_colors,
node_size=3000, font_size=9, connectionstyle="arc3,rad=0.2"
)
nx.draw_networkx_edge_labels(G, pos, edge_labels=edges_labels, font_size=8)
plt.title("OPRAH™ aAPCs", fontsize=18)
plt.show()
# Run the visualization
visualize_nn()
#
Fig. 30 Musical Grammar & Prosody. From a pianist’s perspective, the left hand serves as the foundational architect, voicing the mode and defining the musical landscape—its space and grammar—while the right hand acts as the expressive wanderer, freely extending and altering these modal terrains within temporal pockets, guided by prosody and cadence. In R&B, this interplay often manifests through rich harmonic extensions like 9ths, 11ths, and 13ths, with chromatic passing chords and leading tones adding tension and color. Music’s evocative power lies in its ability to transmit information through a primal, pattern-recognizing architecture, compelling listeners to classify what they hear as either nurturing or threatening—feeding and breeding or fight and flight. This makes music a high-risk, high-reward endeavor, where success hinges on navigating the fine line between coherence and error. Similarly, pattern recognition extends to literature, as seen in Ulysses, where a character misinterprets his companion’s silence as mental composition, reflecting on the instructive pleasures of Shakespearean works used to solve life’s complexities. Both music and literature, then, are deeply rooted in the human impulse to decode and derive meaning, whether through harmonic landscapes or textual introspection. Source: Ulysses, DeepSeek & Yours Truly!#

