Probability, ∂Y 🎶 ✨ 👑#
+ Expand
- The first portrait of Gen. Milley, from his time as the U.S. military's top officer, was removed from the Pentagon last week on Inauguration Day less than two hours after President Trump was sworn into office.
- The now retired Gen. Milley and other former senior Trump aides had been assigned personal security details ever since Iran vowed revenge for the killing of Qasem Soleimani in a drone strike in 2020 ordered by Trump in his first term.
- On "Fox News Sunday," the chairman of the Senate Intelligence Committee, Tom Cotton, said he hoped President Trump would "revisit" the decision to pull the protective security details from John Bolton, Mike Pompeo, and Brian Hook who previously served under Trump.
- A senior administration official who requested anonymity replied, "There is a new era of accountability in the Defense Department under President Trump's leadership—and that's exactly what the American people expect."
- Gen. Milley served as chairman of the Joint Chiefs of Staff from 2019 to 2023 under both Presidents Trump and Biden.
-- Fox News
🪙 🎲 🎰 🐜 🗡️ 🪖 🛡️#
The model begins with the premise that the ecosystem is to be navigated, not imposed upon. At a 95/5 noise-to-signal ratio, the sheer expanse of unknowns, contingencies, and shifting equilibria dwarfs any singular interpretation of order. The world does not conform to preordained structures; it is an ocean of variables in which the agent is a mere speck. To navigate it is not to dictate its terms but to extract meaningful paths through an overwhelming landscape of noise. This is the foundation upon which intelligence must function—not as an autonomous architect but as an adaptive system that filters chaos into coherence. The ecosystem provides the combinatorial search space, and intelligence is, at best, a process of tracking transient footholds within it.
PRR Type |
Receptor Family |
Ligands Recognized |
Location |
Key Functions |
|---|---|---|---|---|
Scavenger Receptors (SRs) |
SR-A, CD36 |
Modified LDL, apoptotic cells, microbial components |
Cell membrane |
Phagocytosis, lipid metabolism, immune clearance |
Fc Receptors (FcRs) |
FcγR, FcεR |
Antibody-antigen complexes |
Cell membrane |
Antibody-dependent immune activation, phagocytosis |
C-type Lectin Receptors (CLRs) |
CLRs (e.g., Dectin-1, DC-SIGN) |
Carbohydrates from fungi, bacteria, and viruses |
Cell membrane |
Fungal recognition, phagocytosis, cytokine modulation |
Toll-like Receptors (TLRs) |
TLRs (e.g., TLR1-TLR10) |
PAMPs (LPS, dsRNA, flagellin) |
Cell membrane & endosomes |
Innate immune activation, cytokine production |
NOD-like Receptors (NLRs) |
NLRs (e.g., NOD1, NOD2, NLRP3) |
Bacterial peptidoglycan, DAMPs |
Cytoplasm |
Intracellular pathogen sensing, inflammasome activation |
RIG-I-like Receptors (RLRs) |
RLRs (e.g., RIG-I, MDA5) |
Viral RNA (dsRNA, ssRNA) |
Cytoplasm |
Antiviral immune response, IFN production |
AIM2-like Receptors (ALRs) |
AIM2, IFI16 |
Cytoplasmic DNA (viral, bacterial, self-DNA) |
Cytoplasm |
Inflammasome activation, IL-1β production |

Fig. 1 Digital Library. Our color-coded QR code library with a franchize model for the digital twin will be launched in a month. The books will explore struggle, exchange, and consolidation as dynamic equilibria that emerge from the oscillation and rhythm of existence.#
Important
Ecosystem 🔐 : Unlocking
Key 🔑: QR code
Agency ℹ️: Learning, Teaching, Information
Wisdom 📖: Past, Treasures
Intelligence 🧞♂️: Dynamic, Grounded
🎭#
Pattern recognition, hardcoded at 80/20, emerges as the first line of filtration. This is where noise is first tempered into partial structure. The sensory ganglia, G1 and G2, function as gatekeepers, automatically encoding patterns that have evolutionary weight. These are not learned truths but precomputed heuristics, the scaffolding upon which later cognition will be built. The high noise ratio here acknowledges that even within these pre-hardwired processes, ambiguity persists—hence, the 20% allowance for correction. This suggests that what appears to be innate is itself an iterative product of past iterations, a record of what has worked but not necessarily what will continue to work. Intelligence does not emerge ex nihilo; it refines inherited biases, selectively overriding static interpretations when the noise-to-signal ratio demands it.
Feature |
MHC (Major Histocompatibility Complex) |
PRRs (Pattern Recognition Receptors) |
|---|---|---|
Type of Immunity |
Adaptive immunity (T-cell activation) |
Innate immunity (pathogen sensing) |
Function |
Presents antigens to T cells |
Detects conserved molecular patterns of pathogens |
Molecular Class |
Polymorphic surface glycoproteins |
Germline-encoded pattern recognition proteins |
Ligands |
Peptides derived from pathogens or self-proteins |
Pathogen-Associated Molecular Patterns (PAMPs) & Damage-Associated Molecular Patterns (DAMPs) |
Cellular Expression |
MHC-I: All nucleated cells; MHC-II: Professional APCs (dendritic cells, \(\text{m}\Phi\), B cells) |
Primarily innate immune cells (\(\text{m}\Phi\), dendritic cells, neutrophils) |
Pathogen Response |
Indirect—activates T cells for specific immunity |
Direct—triggers immediate immune responses (inflammation, phagocytosis) |

Fig. 2 Musk Off. The genome encodes self, other, and mutual interactions within the ecosystem, while the exposome shapes our immune interface through sentinel PRRs, MHC-II, MHC-I, and CD3-ILCs. When these interact with the TCR, transcriptomic shifts drive clonal expansion, triggering a proteasome-mediated cytokine storm from both innate and adaptive systems, ultimately directing the metabolome to mount a robust yet balanced immune response. NK cells (the best known ILC subtype) accounts for 5-20% of circulating lymphocytes. Somatic mutations have the potential to encode “non-self” immunogenic antigens. We hypothesized that tumors with a large number of somatic mutations due to mismatch-repair defects may be susceptible to immune checkpoint blockade. Source: Dung T. Le et al#
🌊 🏄🏾#
At the learned level, fixed versus dynamic processes engage in a 50/50 struggle, each vying for epistemic authority. Fixed learning relies on stored priors, memory encoded through descending fibers, while dynamic learning operates on live input, a function of sensory-driven thalamocortical processing. The equilibrium here is crucial. Lean too heavily on the fixed, and the system stagnates, favoring preemptive conclusions over adaptation. Lean too heavily on the dynamic, and the system becomes noise-chasing, mistaking every transient input for a meaningful shift in structure. This is the pivot upon which intelligence either ossifies or remains fluid. The role of cerebellar-pontine integration, in this sense, is not just computational but existential—determining whether a being remains bound to the past or remains in play within the unfolding uncertainty of the present.
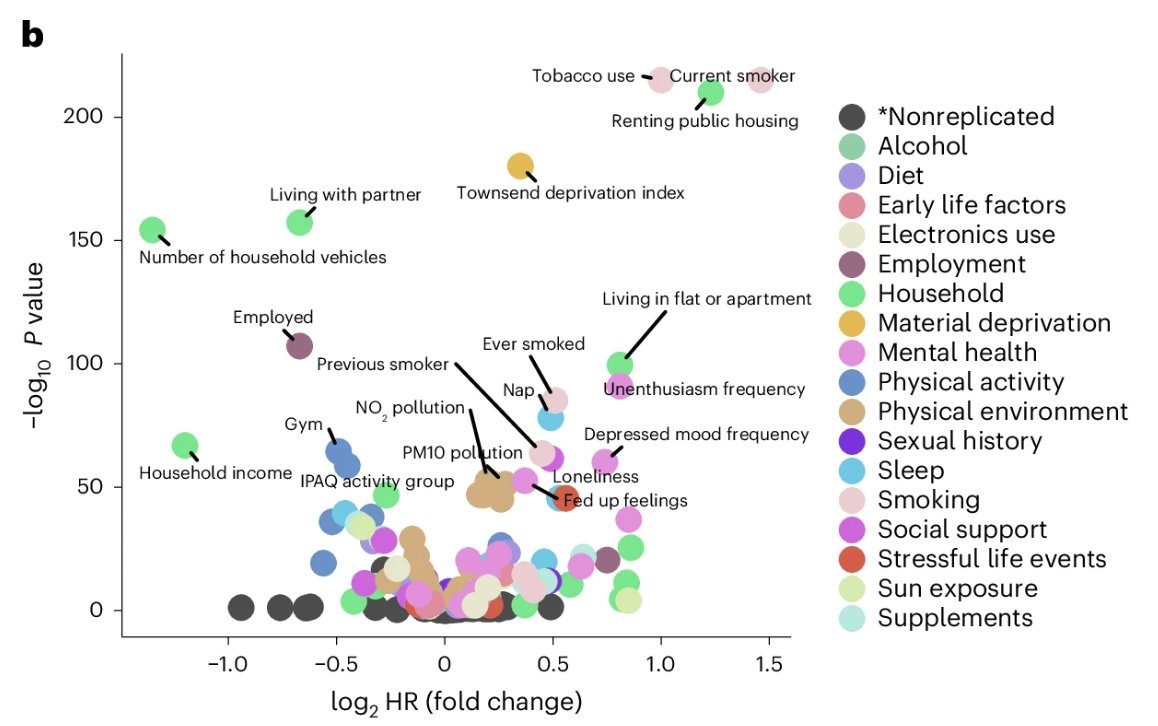
Fig. 3 Lifestyle is a stronger predictor of aging and mortality than genetics. A new Nature study of ~500,000 people found: 🎯 17% of mortality variation was linked to lifestyle (exposome 🚬 🥃 ). 🧬 <2% of was explained by genetics. Here’s what you need to know ☝🏾 🧵 1/10#
🤺 💵 🦘#
The classification of discoveries within the ecosystem follows the triadic structure of Other, Mutual, and Self, mapped onto a 20/80 noise-to-signal ratio. This is where Kantian categorical imperatives find their footing. The moral architecture of intelligence emerges not from the void but from the necessity of sorting relational structures within the combinatorial search space. The Self, dominant at 80%, operates as the primary locus of computation, while the Other and the Mutual, at 20%, serve as correctives. This is an admission that intelligence, by default, is egocentric. The raw signal of survival demands that the self take primacy. However, the 20% remainder forces engagement with externality, ensuring that intelligence does not collapse into solipsism. The Kantian imperative arises here as a tension: if the model is to extend beyond immediate survival, it must recognize the validity of the Other, transitioning from purely adversarial play to cooperative and transactional equilibria.
Danger
Self (cooperative)
Nonself (adversarial)
Mismatch-Repair (transactional)
Cited by 10200 by 10th Anniversary
– by DT Le 2015
🏇 🧘🏾♀️ 🪺 🎶 🛌#
Navigation, at 5/95, represents the final and most precarious stage. It is here that hubris arises—the belief that what is static and structured is all that matters, that self-generated order has subdued the ecosystem’s entropy. But this is a false finality. What appears as a dominant equilibrium—the Ubermensch who believes they have conquered all—is itself a momentary illusion. The 5% noise in navigation can metastasize into something that ripples across the ecosystem, exposing vulnerability when aging and frailty emerge-thus a system must never mistake fixed coordinates for immutable truths. The Ubermensch will be usurped, not because of moral failing, but because the ecosystem does not tolerate stagnation, the Red Queen reigns in perpetuity throughout the universe and you never step in the same river twice. All intelligence is a momentary scaffold upon which future iterations will stand, only to be dismantled again. The true lesson of navigation is not mastery but the inevitability of recalibration.
Show code cell source
import numpy as np
import matplotlib.pyplot as plt
import networkx as nx
# Define the neural network layers
def define_layers():
return {
'Suis': ['DNA, RNA, 5%', 'Peptidoglycans, Lipoteichoics', 'Lipopolysaccharide', 'N-Formylmethionine', "Glucans, Chitin", 'Specific Antigens'],
'Voir': ['PRR & ILCs, 20%'],
'Choisis': ['CD8+, 50%', 'CD4+'],
'Deviens': ['TNF-α, IL-6, IFN-γ', 'PD-1 & CTLA-4', 'Tregs, IL-10, TGF-β, 20%'],
"M'èléve": ['Complement System', 'Platelet System', 'Granulocyte System', 'Innate Lymphoid Cells, 5%', 'Adaptive Lymphoid Cells']
}
# Assign colors to nodes
def assign_colors():
color_map = {
'yellow': ['PRR & ILCs, 20%'],
'paleturquoise': ['Specific Antigens', 'CD4+', 'Tregs, IL-10, TGF-β, 20%', 'Adaptive Lymphoid Cells'],
'lightgreen': ["Glucans, Chitin", 'PD-1 & CTLA-4', 'Platelet System', 'Innate Lymphoid Cells, 5%', 'Granulocyte System'],
'lightsalmon': ['Lipopolysaccharide', 'N-Formylmethionine', 'CD8+, 50%', 'TNF-α, IL-6, IFN-γ', 'Complement System'],
}
return {node: color for color, nodes in color_map.items() for node in nodes}
# Define edge weights
def define_edges():
return {
('DNA, RNA, 5%', 'PRR & ILCs, 20%'): '1/99',
('Peptidoglycans, Lipoteichoics', 'PRR & ILCs, 20%'): '5/95',
('Lipopolysaccharide', 'PRR & ILCs, 20%'): '20/80',
('N-Formylmethionine', 'PRR & ILCs, 20%'): '51/49',
("Glucans, Chitin", 'PRR & ILCs, 20%'): '80/20',
('Specific Antigens', 'PRR & ILCs, 20%'): '95/5',
('PRR & ILCs, 20%', 'CD8+, 50%'): '20/80',
('PRR & ILCs, 20%', 'CD4+'): '80/20',
('CD8+, 50%', 'TNF-α, IL-6, IFN-γ'): '49/51',
('CD8+, 50%', 'PD-1 & CTLA-4'): '80/20',
('CD8+, 50%', 'Tregs, IL-10, TGF-β, 20%'): '95/5',
('CD4+', 'TNF-α, IL-6, IFN-γ'): '5/95',
('CD4+', 'PD-1 & CTLA-4'): '20/80',
('CD4+', 'Tregs, IL-10, TGF-β, 20%'): '51/49',
('TNF-α, IL-6, IFN-γ', 'Complement System'): '80/20',
('TNF-α, IL-6, IFN-γ', 'Platelet System'): '85/15',
('TNF-α, IL-6, IFN-γ', 'Granulocyte System'): '90/10',
('TNF-α, IL-6, IFN-γ', 'Innate Lymphoid Cells, 5%'): '95/5',
('TNF-α, IL-6, IFN-γ', 'Adaptive Lymphoid Cells'): '99/1',
('PD-1 & CTLA-4', 'Complement System'): '1/9',
('PD-1 & CTLA-4', 'Platelet System'): '1/8',
('PD-1 & CTLA-4', 'Granulocyte System'): '1/7',
('PD-1 & CTLA-4', 'Innate Lymphoid Cells, 5%'): '1/6',
('PD-1 & CTLA-4', 'Adaptive Lymphoid Cells'): '1/5',
('Tregs, IL-10, TGF-β, 20%', 'Complement System'): '1/99',
('Tregs, IL-10, TGF-β, 20%', 'Platelet System'): '5/95',
('Tregs, IL-10, TGF-β, 20%', 'Granulocyte System'): '10/90',
('Tregs, IL-10, TGF-β, 20%', 'Innate Lymphoid Cells, 5%'): '15/85',
('Tregs, IL-10, TGF-β, 20%', 'Adaptive Lymphoid Cells'): '20/80'
}
# Define edges to be highlighted in black
def define_black_edges():
return {
('DNA, RNA, 5%', 'PRR & ILCs, 20%'): '1/99',
('Peptidoglycans, Lipoteichoics', 'PRR & ILCs, 20%'): '5/95',
('Lipopolysaccharide', 'PRR & ILCs, 20%'): '20/80',
('N-Formylmethionine', 'PRR & ILCs, 20%'): '51/49',
("Glucans, Chitin", 'PRR & ILCs, 20%'): '80/20',
('Specific Antigens', 'PRR & ILCs, 20%'): '95/5',
}
# Calculate node positions
def calculate_positions(layer, x_offset):
y_positions = np.linspace(-len(layer) / 2, len(layer) / 2, len(layer))
return [(x_offset, y) for y in y_positions]
# Create and visualize the neural network graph
def visualize_nn():
layers = define_layers()
colors = assign_colors()
edges = define_edges()
black_edges = define_black_edges()
G = nx.DiGraph()
pos = {}
node_colors = []
# Create mapping from original node names to numbered labels
mapping = {}
counter = 1
for layer in layers.values():
for node in layer:
mapping[node] = f"{counter}. {node}"
counter += 1
# Add nodes with new numbered labels and assign positions
for i, (layer_name, nodes) in enumerate(layers.items()):
positions = calculate_positions(nodes, x_offset=i * 2)
for node, position in zip(nodes, positions):
new_node = mapping[node]
G.add_node(new_node, layer=layer_name)
pos[new_node] = position
node_colors.append(colors.get(node, 'lightgray'))
# Add edges with updated node labels
edge_colors = []
for (source, target), weight in edges.items():
if source in mapping and target in mapping:
new_source = mapping[source]
new_target = mapping[target]
G.add_edge(new_source, new_target, weight=weight)
edge_colors.append('black' if (source, target) in black_edges else 'lightgrey')
# Draw the graph
plt.figure(figsize=(12, 8))
edges_labels = {(u, v): d["weight"] for u, v, d in G.edges(data=True)}
nx.draw(
G, pos, with_labels=True, node_color=node_colors, edge_color=edge_colors,
node_size=3000, font_size=9, connectionstyle="arc3,rad=0.2"
)
nx.draw_networkx_edge_labels(G, pos, edge_labels=edges_labels, font_size=8)
plt.title("OPRAH™ & ARNTC", fontsize=18)
plt.show()
# Run the visualization
visualize_nn()


Fig. 4 Transactional Imperative of DNA Mismatch Repair: PD-1 is a Token for Continued Dealing. MMR is a system for recognizing and repairing erroneous insertion, deletion, and mis-incorporation of bases that can arise during DNA replication and recombination, as well as repairing some forms of DNA damage. Mismatch repair is strand-specific. During DNA synthesis the newly synthesised (daughter) strand will commonly include errors. In order to begin repair, the mismatch repair machinery distinguishes the newly synthesised strand from the template (parental). In gram-negative bacteria, transient hemimethylation distinguishes the strands (the parental is methylated and daughter is not). However, in other prokaryotes and eukaryotes, the exact mechanism is not clear. It is suspected that, in eukaryotes, newly synthesized lagging-strand DNA transiently contains nicks (before being sealed by DNA ligase) and provides a signal that directs mismatch proofreading systems to the appropriate strand. This implies that these nicks must be present in the leading strand, and evidence for this has recently been found. Recent work has shown that nicks are sites for RFC-dependent loading of the replication sliding clamp, proliferating cell nuclear antigen (PCNA), in an orientation-specific manner, such that one face of the donut-shape protein is juxtaposed toward the 3’-OH end at the nick. Loaded PCNA then directs the action of the MutLalpha endonuclease to the daughter strand in the presence of a mismatch and MutSalpha or MutSbeta.#
The immune system operates as a complex and layered network of surveillance, recognition, and response mechanisms, structured to maintain homeostasis while distinguishing between self and non-self. At its foundation, the genome encodes the intrinsic blueprint of immunity, categorizing interactions within the ecosystem into self, other, and mutual elements. The exposome—comprising environmental exposures—modulates immune recognition through key sentinels such as Pattern Recognition Receptors (PRRs), Major Histocompatibility Complex (MHC) molecules, and innate lymphoid cells. PRRs, including Toll-like receptors (TLRs), detect conserved microbial patterns, while MHC molecules coordinate antigen presentation. Class I MHC primarily interacts with CD8+ cytotoxic T cells, surveilling intracellular infections, whereas Class II MHC engages CD4+ helper T cells, orchestrating adaptive immunity against extracellular threats. These molecular interactions initiate a cascade of immune responses, mediated through transcriptomic shifts and clonal expansion of T cells following engagement with the T-cell receptor (TCR).
Upon TCR activation, intracellular signaling pathways trigger a proteasome-mediated degradation of foreign peptides, followed by cytokine production to refine and direct immune activity. Pro-inflammatory cytokines such as TNF-α, IL-6, and IFN-γ amplify innate defenses, while regulatory molecules, including PD-1 and CTLA-4, temper excessive activation to prevent autoimmunity. The metabolome—the biochemical output of immune signaling—ultimately integrates these signals to fine-tune physiological responses. This dynamic interplay between innate and adaptive immunity ensures a robust yet controlled defense, preserving host integrity while adapting to evolving microbial threats.
Mapping this immunological framework onto a neural network architecture offers an intriguing model for conceptualizing information processing in biological systems. Both immune and neural networks rely on layers of signal transduction, feedback mechanisms, and adaptability to maintain equilibrium. However, the analogy breaks down when considering the fundamental distinctions between the two systems. Neural networks operate through electrochemical signaling with a relatively fixed wiring pattern and continuous feedback loops, whereas immune responses rely on stochastic gene recombination, clonal selection, and transient cellular interactions. While neural circuits emphasize precision and hierarchical decision-making, immune signaling involves probabilistic encounters, dynamic plasticity, and decentralized control. Thus, while the immune system shares computational parallels with neural networks in terms of pattern recognition and adaptive responses, its stochastic, population-based decision-making diverges from the structured, gradient-based optimization of artificial intelligence models. Any attempt to superimpose neural architecture onto immune dynamics must account for these fundamental disparities in information processing and evolutionary constraints.

